
Using skrmdb
Thomas Kent
2026-01-16
using_skrmdb.rmdNotation
The methods of Spearman-Karber, Reed-Muench, and Dragstedt-Behrens are all commonly used to estimate ED50. In what follows, we use the following variable notation:
-
xis a vector corresponding to the log dilution or dose for each group. -
nis an integer vector corresponding to the group size at each log dilution or dose. -
yis an integer vector corresponding to the number responding at each log dilution or dose.
All examples are with SpearKarb(), however, the usage
for DragBehr() and ReedMuench() is
identical.
Usage
Each of the main functions in skrmdb can be called in
three different ways.
To illustrate this, we start with a simple data set where the number of dead increases with the dosage.
First we use the historical, deprecated, function call.
SpearKarb(y = dead, n = total, x = dil)## Warning: skrmdb :: Calling this function with the parameters y, n, x is
## depreciated.## ED50 by the Spearman-Karber method
## ed: 2.9## Count trend is increasing with dilution.The preferred method is to use formulas in the function call.
We can use the formula on the individual vectors.
SpearKarb(dead + total ~ dil)## ED50 by the Spearman-Karber method
## ed: 2.9## Count trend is increasing with dilution.We can also use the formula on columns in a data.frame.
data <- data.frame(y = dead, n = total, x = dil)
SpearKarb(y + n ~ x, data)## ED50 by the Spearman-Karber method
## ed: 2.9## Count trend is increasing with dilution.Method Assumptions
Each of these methods was designed to work under the assumption that x is either increasing or decreasing and y / n is increasing with the index.
It was decided that the functions DragBehr(),
ReedMuench(), and SpearKarb() will sort the
data for you according to this assumption.
To illustrate this, we create some data that is descending according to dilution.
And we see that no mater how we enter the data, the same ED50 is reported.
SpearKarb(dead + total ~ dil)## ED50 by the Spearman-Karber method
## ed: 4.1## Count trend is decreasing with dilution.## ED50 by the Spearman-Karber method
## ed: 4.1## Count trend is decreasing with dilution.However, if we can decided to turn off the autosort, which will give us a different, but incorrect, estimate of ED50.
SpearKarb(dead + total ~ dil, autosort = FALSE)## ED50 by the Spearman-Karber method
## ed: 2.9## Count trend is decreasing with dilution.Conditional ED50
The function skrmdb.all returns the ED50 for the three
methods, along with additional data about the data sets that may be of
interest.
For this section, we use the example data set titration.
This data.frame contains the results of a hypothetical
experiment where the ED50 of three vials (numbered 1, 2, 3) were each
tested by three anonymous operators (TK, NU, CT).
head(titration)## testID PrepID PrepRole Date Vial Operator dil positive total
## 1 BRSV BRSV-001 test 03-Mar-19 1 TK 1e-02 10 10
## 2 BRSV BRSV-001 test 03-Mar-19 1 TK 1e-03 10 10
## 3 BRSV BRSV-001 test 03-Mar-19 1 TK 1e-04 9 10
## 4 BRSV BRSV-001 test 03-Mar-19 1 TK 1e-05 6 10
## 5 BRSV BRSV-001 test 03-Mar-19 1 TK 1e-06 1 10
## 6 BRSV BRSV-001 test 03-Mar-19 1 TK 1e-07 1 10First we need to compute the log dilution.
titration$log_dil <- -log10(titration$dil)Plotting the data shows us that the tests visually seem to give roughly the same ED50, but unfortunately, there are missing data points.
ggplot(titration, aes(x = log_dil, y = positive)) +
geom_point() +
facet_grid(Vial ~ Operator) +
geom_line()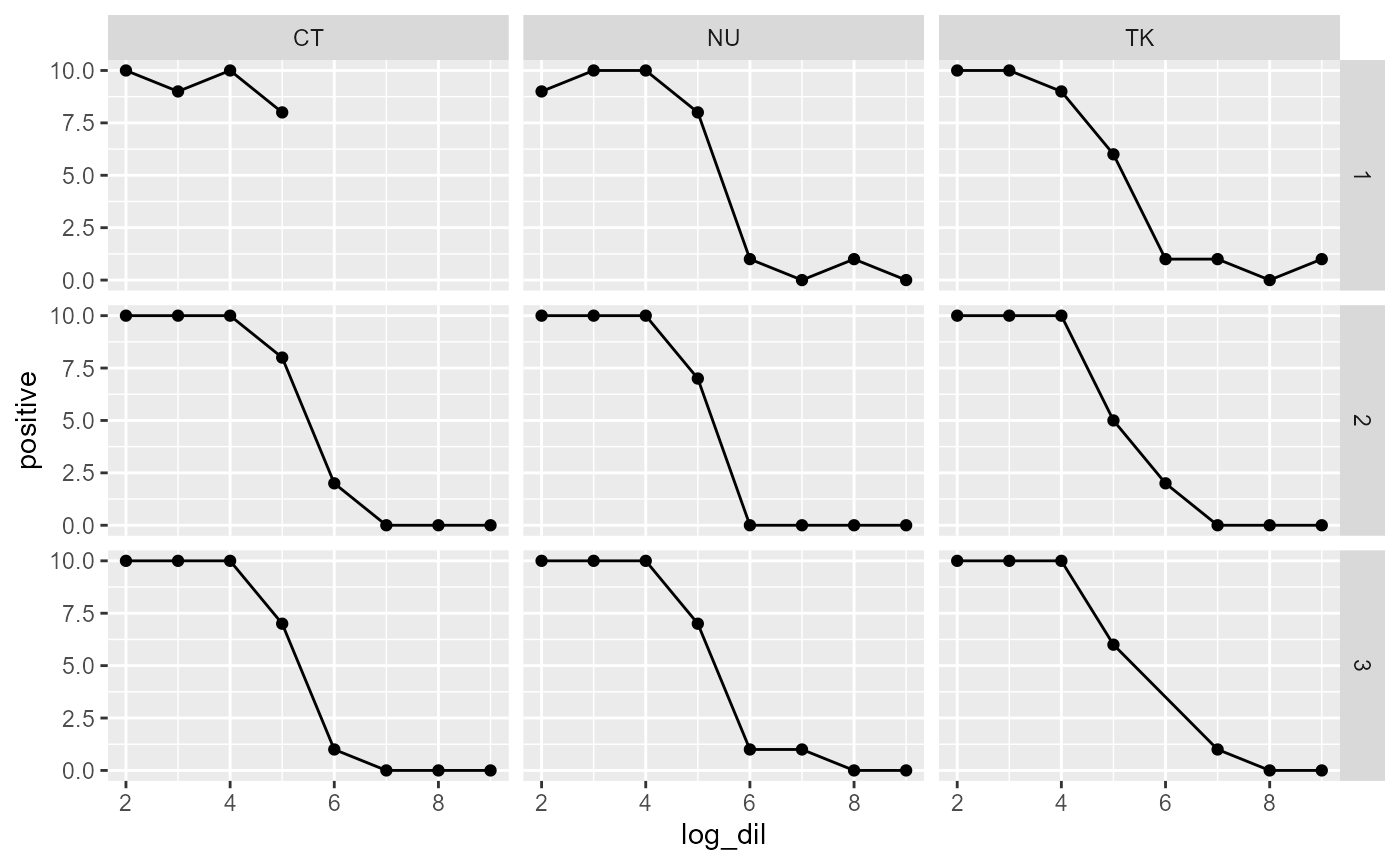
We could find ED50 by aggregation the data from all 9 tests for each of the three methods.
skrmdb.all(positive + total ~ log_dil, titration)## DragBehr ReedMuench SpearKarb SpearKarb.var response.increasing
## 1 5.365201 5.331023 5.332341 0.005014876 FALSE
## duplicate.dilutions even.dilution monotonic bracket.midpoint
## 1 TRUE TRUE TRUE TRUEWe can also see if there is a difference between Operators;
skrmdb.all(positive + total ~ log_dil | Operator, titration)## Operator DragBehr ReedMuench SpearKarb SpearKarb.var response.increasing
## 1 TK 5.286004 5.247059 5.283333 0.01954578 FALSE
## 2 NU 5.376350 5.340000 5.333333 0.01222222 FALSE
## 3 CT 5.418526 5.402062 5.383333 0.01399022 FALSE
## duplicate.dilutions even.dilution monotonic bracket.midpoint
## 1 TRUE TRUE FALSE TRUE
## 2 TRUE TRUE FALSE TRUE
## 3 TRUE TRUE FALSE TRUEOr by Vial;
skrmdb.all(positive + total ~ log_dil | Vial, titration)## Vial DragBehr ReedMuench SpearKarb SpearKarb.var response.increasing
## 1 1 5.405272 5.377551 5.383333 0.02231347 FALSE
## 2 2 5.339806 5.304348 5.300000 0.01164751 FALSE
## 3 3 5.360000 5.319149 5.333333 0.01454527 FALSE
## duplicate.dilutions even.dilution monotonic bracket.midpoint
## 1 TRUE TRUE TRUE TRUE
## 2 TRUE TRUE TRUE TRUE
## 3 TRUE TRUE TRUE TRUEOr by Operator and Vial.
skrmdb.all(positive + total ~ log_dil | Vial + Operator, titration)## Vial Operator DragBehr ReedMuench SpearKarb SpearKarb.var response.increasing
## 1 1 TK 5.306306 5.266667 5.30 0.06666667 FALSE
## 2 1 NU 5.429825 5.411765 5.40 0.04777778 FALSE
## 3 1 CT NA NA 5.20 0.02777778 FALSE
## 4 2 TK 5.185185 5.153846 5.20 0.04555556 FALSE
## 5 2 NU 5.285714 5.235294 5.20 0.02333333 FALSE
## 6 2 CT 5.500000 5.500000 5.50 0.03555556 FALSE
## 7 3 TK 5.482759 5.400000 5.55 0.08250000 FALSE
## 8 3 NU 5.411765 5.375000 5.40 0.04333333 FALSE
## 9 3 CT 5.349462 5.312500 5.30 0.03333333 FALSE
## duplicate.dilutions even.dilution monotonic bracket.midpoint
## 1 FALSE TRUE FALSE TRUE
## 2 FALSE TRUE FALSE TRUE
## 3 FALSE TRUE FALSE FALSE
## 4 FALSE TRUE TRUE TRUE
## 5 FALSE TRUE TRUE TRUE
## 6 FALSE TRUE TRUE TRUE
## 7 FALSE FALSE TRUE TRUE
## 8 FALSE TRUE TRUE TRUE
## 9 FALSE TRUE TRUE TRUEAdditional Methods
Finally, there are three accessor functions to help retrieve
information from the results of DragBehr(),
ReedMuench(), and SpearKarb().
Using the titration data set again.
res <- SpearKarb(positive + total ~ log_dil, titration)## skrmdb :: combining results from duplicate dilutionsWe can get the ED50: (Note: getED50()
only works on skrmdb objects, which are returned by
DragBehr(), ReedMuench(), and
SpearKarb(), not on skrmdb.all objects.)
getED50(res)## [1] 5.332341The variance:
getvar(res)## [1] 0.005014876And the data which was used.
getdata(res)## x y n y_inc y_dec
## 1 2 89 90 5600 498400
## 2 3 89 90 5600 498400
## 3 4 89 90 5600 498400
## 4 5 62 90 156800 347200
## 5 6 8 70 446400 57600
## 6 7 3 80 485100 18900
## 7 8 1 80 497700 6300
## 8 9 1 80 497700 6300